Quantitative descriptions of biomolecular dynamics: computational and spectroscopic descriptors
Thermal motions of biomolecules drive many cellular processes crucial for life. However, the specific links between protein dynamics and function remain unclear at present. In fact, even the extent to which protein dynamics participates in biomolecular reactions is a topic of passionate debate between prominent chemists. The advent of computational modelling and simulation techniques and developments in computer hardware technology have enabled high resolution atomistic descriptions of biomolecular dynamics over microsecond-millisecond timescales.Nevertheless, dynamical descriptors for biomolecular function have remained elusive as they lie embedded in a complex multi-dimensional space of soft degrees of freedom which span an impressive 15 to 18 orders of magnitude ranging from localized bond vibration motions on the femtosecond timescale to protein translocation on the seconds to hours timescale. In general, extracting functionally relevant dynamical biomolecular descriptors is a formidable unsolved challenge for both theory and experiment. Our group has made progress in this area by developing computational and spectroscopic descriptors of biomolecular dynamics which can be linked to protein function.
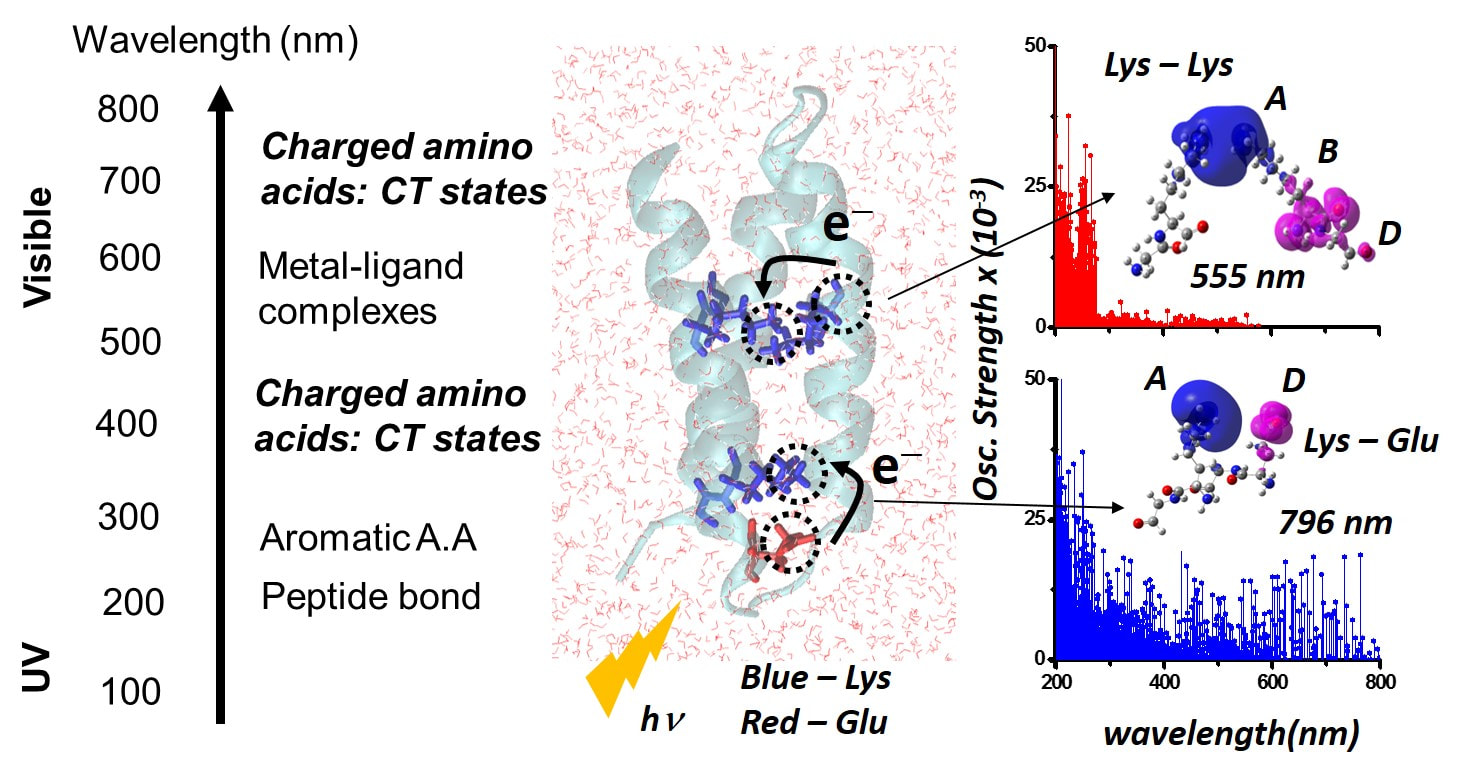
Electronic Charge Transfer Bands to Probe Biomolecular Structure and DynamicsOptical spectroscopy provides a powerful means to examine the chemical composition, structure, and dynamics of biomolecules. The most prominent features (Figure: top right) in the UV-Visible spectra of proteins arise (Figure on the right) from the peptide backbone, three dominant aromatic amino acid chromophores (tryptophan, tyrosine, and phenylalanine), and .specialized metal containing or aromatic co-factors at protein active-sites. As such, the electronic absorption properties of the majority of natural amino acids in proteins remain untapped at present, and proteins lacking active site chromophores or aromatic amino acids are expected to remain optically silent above 300 nm. We have discovered distinctive mechanisms of photoinduced charge transfer (CT) transitions arising from charged amino acids (lysine, glutamate, histidine, arginine, and aspartate) in the UV-Visible region of the electromagnetic spectrum for proteins (Figure: top left).
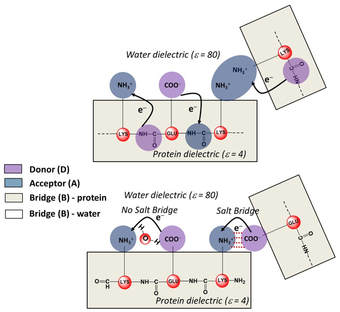
The general principle uncovered by us is that charged amino acids, in isolation or collective assemblies, form electronic Donor(D) – Bridge(B) – Acceptor(A) molecular complexes which exhibit photoinduced D to A CT transitions (Figure: bottom right). The charged sidechain moieties, such as lysine amino and glutamate carboxylate groups, and the peptide backbone play the role of D and A groups, while the aliphatic part of the sidechain or the intervening protein/solvent medium form the B component of the molecular complex (Figure below). By establishing the first quantitative link between non-aromatic protein amino acid sequence content and the UV-Vis absorption features above 320 nm, we have opened up a new spectral window to probe structure, dynamics, and function of prominent proteins of biomedical relevance. |
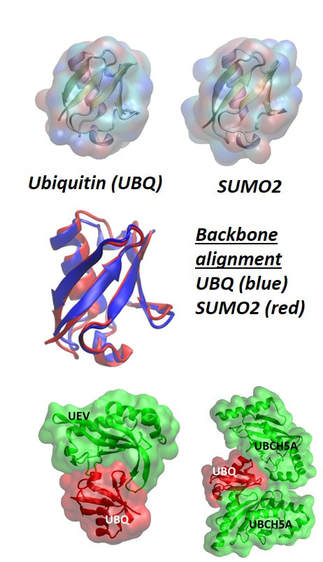
Sorting the Function of Ubiquitin Family ProteinsProteins from the ubiquitin family, which include ubiquitin and small ubiquitin-like modifier (SUMO) proteins, serve as molecular tags that annotate proteins for many different cellular functions necessary for life, ranging from protein degradation to DNA transcription, replication, and repair. The ubiquitin and SUMO proteins share a strong structural homology exhibiting a superimposable b-grasp fold (Figure:on the left). The general mechanism of activation and attachment of ubiquitin and SUMO tags to proteins is also the same. Despite the structural and mechanistic similarities of ubiquitin and SUMO proteins, the functional role of each molecular tag is distinct. We have proposed that a molecular level descriptor, the equilibrium protein flexibility, can be used as a control parameter by the cellular machinery to activate, assign, and recognize ubiquitin family molecular tags.
Algorithms for Quantitative Analysis of Protein Structures
The protein databank (www.rcsb.org)21 is a treasure-trove of information containing more than 105 X-ray protein structures. Potentially, information about structural changes induced in proteins by binding to other proteins or ligands can be derived by examining multiple structures of the same protein in the databank in complexed and free forms. Our group is developing computational algorithms for quantitative comparison of protein structures from the protein data bank to objectively resolve collective structural changes in proteins induced by the binding of biological and synthetic molecules.Our efforts are aimed at deriving reliable structural information and associated properties of biomolecules directly from experimental data to create models of functional biomolecular motions.
|